Apply ctSVG to Visium HD data
Haotian Zhuang
January 04, 2026
Source:vignettes/visiumHD.Rmd
visiumHD.RmdIntroduction
We will demonstrate how to use ctSVG to generate single-cell gene expression profiles and identify cell-type-specific SVGs from Visium HD data. The Visium HD dataset of a mouse brain tissue was downloaded from the 10x website.
Download Space Ranger output files and H&E image
The Space Ranger output files were downloaded at https://cf.10xgenomics.com/samples/spatial-exp/3.0.0/Visium_HD_Mouse_Brain/Visium_HD_Mouse_Brain_binned_outputs.tar.gz. The H&E image was downloaded at https://cf.10xgenomics.com/samples/spatial-exp/3.0.0/Visium_HD_Mouse_Brain/Visium_HD_Mouse_Brain_tissue_image.tif.
Perform nuclei segmentation
The nuclei segmentation results can be obtained by running segmentation methods, such as StarDist, on the H&E image accompanying the Visium HD data.
import pandas as pd
import numpy as np
from tifffile import imread, imwrite
from csbdeep.utils import normalize
from stardist.models import StarDist2D
img = imread('Visium_HD_Mouse_Brain_tissue_image.tif')
# Load the pretrained model
model = StarDist2D.from_pretrained('2D_versatile_he')
# Percentile normalization of the image
min_percentile = 5
max_percentile = 95
img = normalize(img, min_percentile, max_percentile)
# Stardist
labels, polys = model.predict_instances_big(img, axes='YXC', block_size=4096, prob_thresh=0.01,nms_thresh=0.001, min_overlap=128, context=128, normalizer=None, n_tiles=(4,4,1))
# Convert data format
data = [(cell_id, x, y) for cell_id, (xs, ys) in enumerate(polys['coord']) for x, y in zip(xs, ys)]
df = pd.DataFrame(data, columns=['Cell', 'X', 'Y'])
df.to_csv('nuclei.csv', index=False)Load package
library(ctSVG)
mydir <- "~/Desktop/Research/ctSVG/visiumHD_mouse_brain"
list.files(mydir)
#> [1] "binned_outputs" "nuclei.csv"Assign each square to a cell
suppressMessages(suppressWarnings(
cellassignRes <- cellassign(visiumpath = file.path(mydir, "binned_outputs/square_002um"),
segmentationpath = file.path(mydir, "nuclei.csv"))
))
str(cellassignRes)
#> List of 2
#> $ assign: Named chr [1:1858547] "Cell:21001" "Cell:21001" "Cell:21001" "Cell:21001" ...
#> ..- attr(*, "names")= chr [1:1858547] "s_002um_00029_00776-1" "s_002um_00029_00777-1" "s_002um_00029_00778-1" "s_002um_00029_00779-1" ...
#> $ coord : num [1:82357, 1:2] 355 365 363 364 362 ...
#> ..- attr(*, "dimnames")=List of 2
#> .. ..$ : chr [1:82357] "Cell:21001" "Cell:29526" "Cell:19243" "Cell:31821" ...
#> .. ..$ : chr [1:2] "row" "col"
coord <- cellassignRes$coordGenerate single-cell gene expression profiles
count <- getexpr(cellassignRes = cellassignRes,
visiumpath = file.path(mydir, "binned_outputs/square_002um"))
str(count)
#> Formal class 'dgCMatrix' [package "Matrix"] with 6 slots
#> ..@ i : int [1:33493039] 42 109 198 243 291 364 561 627 744 763 ...
#> ..@ p : int [1:82358] 0 233 265 373 416 436 476 510 623 694 ...
#> ..@ Dim : int [1:2] 19059 82357
#> ..@ Dimnames:List of 2
#> .. ..$ : chr [1:19059] "Xkr4" "Rp1" "Sox17" "Lypla1" ...
#> .. ..$ : chr [1:82357] "Cell:21001" "Cell:29526" "Cell:19243" "Cell:31821" ...
#> ..@ x : num [1:33493039] 1 1 2 1 5 1 1 1 3 1 ...
#> ..@ factors : list()Run Seurat pipeline
suppressMessages(suppressWarnings(
d <- runseurat(count = count)
))
expr <- d@assays$RNA$data
sc <- d$seurat_clusters
str(sc)
#> Factor w/ 26 levels "0","1","2","3",..: 13 13 13 13 11 14 4 3 11 2 ...
#> - attr(*, "names")= chr [1:30558] "Cell:7101" "Cell:5729" "Cell:5749" "Cell:5892" ...Reassign cells to clusters
In this vignette, n.max is set to 200 to reduce
computational time. By default, n.max is set to 1,000 in
practice.
recluRes <- recluster(d = d, n.max = 200)Identify spatially variable genes within each cell cluster
suppressMessages(suppressWarnings(
clutestRes <- ctsvg_test(expr = expr, coord = coord, sc = sc, recluRes = recluRes)
))
head(clutestRes)
#> cluster gene fdr pval fdr.empirical
#> 0.C1ql3 0 C1ql3 1.557701e-293 1.535740e-297 0.05422556
#> 0.Snap25 0 Snap25 1.021168e-212 2.013542e-216 0.05422556
#> 0.Rab3c 0 Rab3c 1.371994e-184 4.057955e-188 0.05422556
#> 0.Nptxr 0 Nptxr 8.103704e-171 3.195782e-174 0.05422556
#> 0.Cyp26b1 0 Cyp26b1 1.441156e-165 7.104190e-169 0.05422556
#> 0.Man1a 0 Man1a 3.105247e-144 1.836881e-147 0.05422556
#> pval.empirical fstat.ori
#> 0.C1ql3 0.00990099 78.98951
#> 0.Snap25 0.00990099 65.73079
#> 0.Rab3c 0.00990099 50.76282
#> 0.Nptxr 0.00990099 46.25440
#> 0.Cyp26b1 0.00990099 79.04013
#> 0.Man1a 0.00990099 41.00661Fit spatial gene expression patterns within each cell cluster
clufitRes <- ctsvg_fit(expr = expr, coord = coord, sc = sc)Visualize spatial patterns of a single gene
plotGene(expr = expr, clufitRes = clufitRes, coord = coord, sc = sc,
gene = clutestRes$gene[1], cellclu = "0", background = TRUE)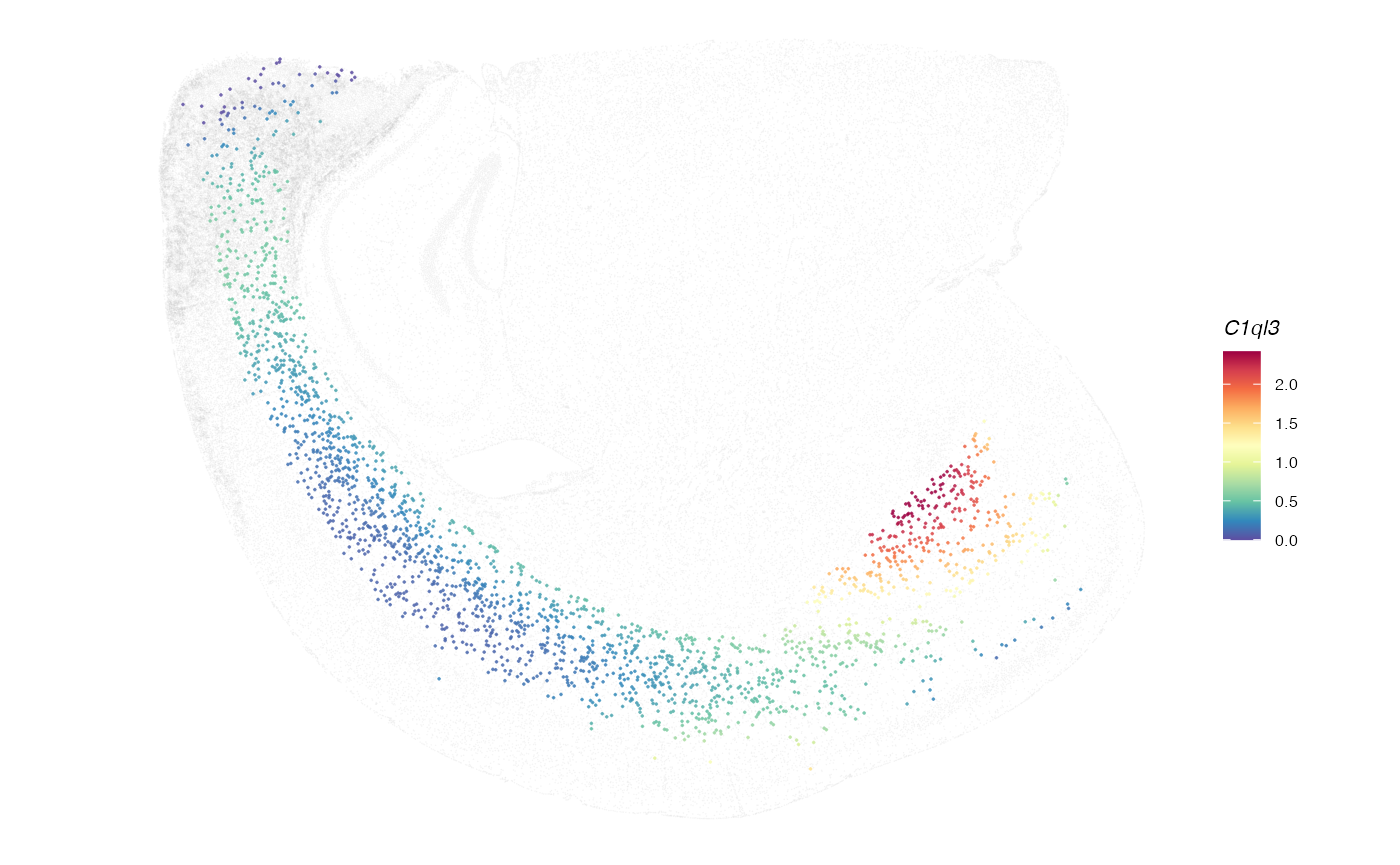
Identify gene modules within each cell cluster
moduleRes <- ctsvg_module(clutestRes = clutestRes, clufitRes = clufitRes, sc = sc)Visualize spatial patterns of metagenes within a cell cluster
plotMetagene(clufitRes = clufitRes, coord = coord, sc = sc,
moduleRes = moduleRes, cellclu = "0", background = TRUE)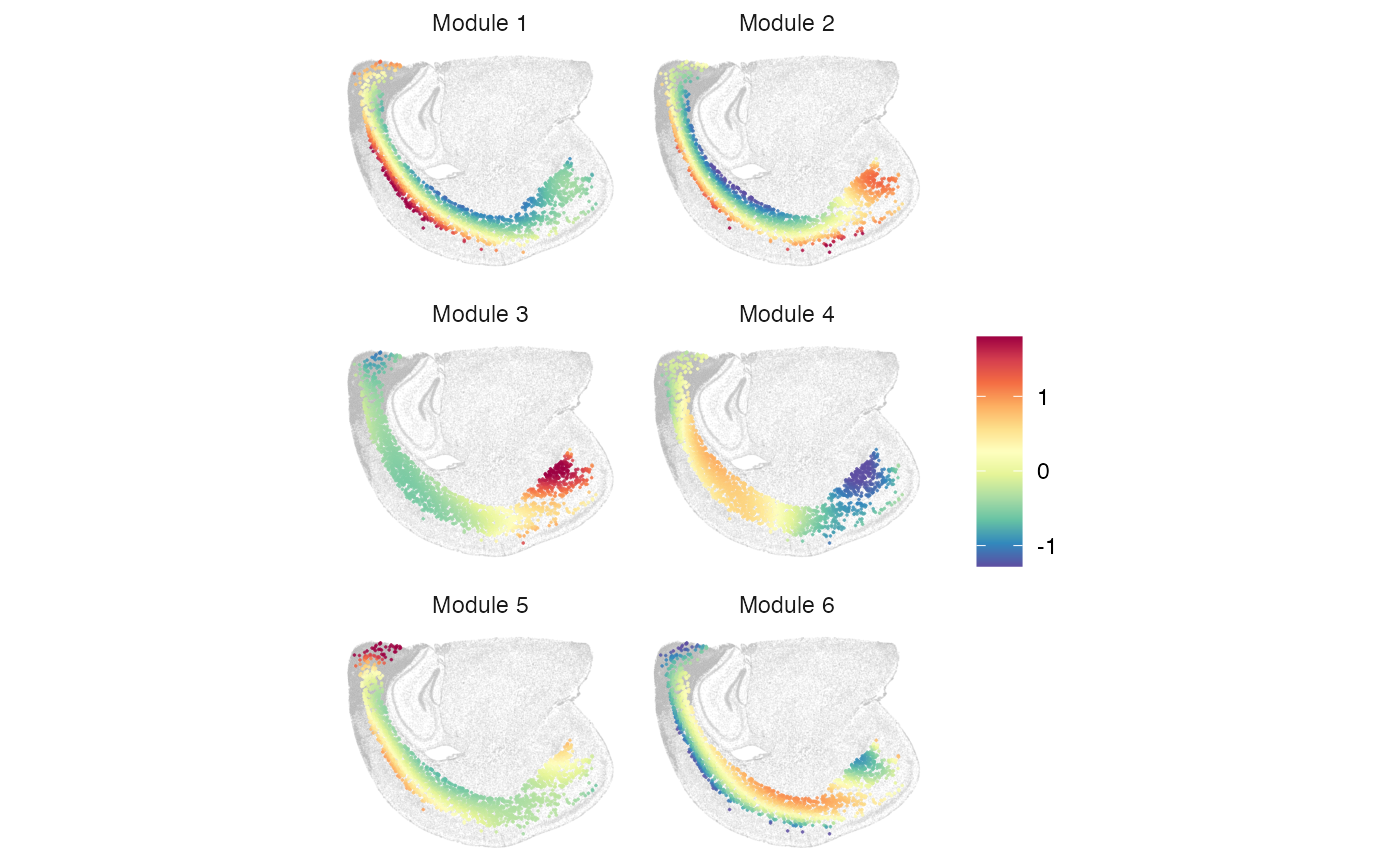
Session Info
sessionInfo()
#> R version 4.4.1 (2024-06-14)
#> Platform: aarch64-apple-darwin20
#> Running under: macOS 15.6.1
#>
#> Matrix products: default
#> BLAS: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRblas.0.dylib
#> LAPACK: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.0
#>
#> locale:
#> [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
#>
#> time zone: America/New_York
#> tzcode source: internal
#>
#> attached base packages:
#> [1] stats graphics grDevices utils datasets methods base
#>
#> other attached packages:
#> [1] ctSVG_1.0
#>
#> loaded via a namespace (and not attached):
#> [1] RColorBrewer_1.1-3 rstudioapi_0.16.0 jsonlite_1.8.8
#> [4] magrittr_2.0.3 spatstat.utils_3.1-0 farver_2.1.2
#> [7] rmarkdown_2.28 zlibbioc_1.50.0 fs_1.6.4
#> [10] ragg_1.3.2 vctrs_0.6.5 ROCR_1.0-11
#> [13] spatstat.explore_3.3-2 S4Arrays_1.4.1 htmltools_0.5.8.1
#> [16] findPC_1.0 SparseArray_1.4.8 sass_0.4.9
#> [19] sctransform_0.4.1 parallelly_1.38.0 KernSmooth_2.23-24
#> [22] bslib_0.8.0 htmlwidgets_1.6.4 desc_1.4.3
#> [25] ica_1.0-3 plyr_1.8.9 plotly_4.10.4
#> [28] zoo_1.8-12 cachem_1.1.0 igraph_2.0.3
#> [31] mime_0.12 lifecycle_1.0.4 pkgconfig_2.0.3
#> [34] Matrix_1.7-0 R6_2.5.1 fastmap_1.2.0
#> [37] MatrixGenerics_1.16.0 fitdistrplus_1.2-1 future_1.34.0
#> [40] shiny_1.9.1 digest_0.6.37 colorspace_2.1-1
#> [43] S4Vectors_0.42.1 patchwork_1.3.0 Seurat_5.1.0
#> [46] tensor_1.5 RSpectra_0.16-2 irlba_2.3.5.1
#> [49] textshaping_0.4.0 labeling_0.4.3 progressr_0.14.0
#> [52] fansi_1.0.6 spatstat.sparse_3.1-0 httr_1.4.7
#> [55] polyclip_1.10-7 abind_1.4-8 compiler_4.4.1
#> [58] proxy_0.4-27 withr_3.0.1 bit64_4.5.2
#> [61] DBI_1.2.3 fastDummies_1.7.4 highr_0.11
#> [64] MASS_7.3-60.2 rjson_0.2.23 classInt_0.4-10
#> [67] units_0.8-5 tools_4.4.1 lmtest_0.9-40
#> [70] httpuv_1.6.15 future.apply_1.11.2 goftest_1.2-3
#> [73] glue_1.7.0 nlme_3.1-164 promises_1.3.0
#> [76] grid_4.4.1 sf_1.0-18 Rtsne_0.17
#> [79] cluster_2.1.6 reshape2_1.4.4 generics_0.1.3
#> [82] hdf5r_1.3.11 gtable_0.3.5 spatstat.data_3.1-2
#> [85] class_7.3-22 tidyr_1.3.1 data.table_1.16.2
#> [88] XVector_0.44.0 sp_2.1-4 utf8_1.2.4
#> [91] BiocGenerics_0.50.0 spatstat.geom_3.3-3 RcppAnnoy_0.0.22
#> [94] ggrepel_0.9.6 RANN_2.6.2 pillar_1.9.0
#> [97] stringr_1.5.1 spam_2.11-0 RcppHNSW_0.6.0
#> [100] later_1.3.2 splines_4.4.1 dplyr_1.1.4
#> [103] lattice_0.22-6 bit_4.5.0 survival_3.6-4
#> [106] deldir_2.0-4 tidyselect_1.2.1 miniUI_0.1.1.1
#> [109] pbapply_1.7-2 knitr_1.48 gridExtra_2.3
#> [112] IRanges_2.38.1 scattermore_1.2 stats4_4.4.1
#> [115] xfun_0.47 matrixStats_1.3.0 stringi_1.8.4
#> [118] lazyeval_0.2.2 yaml_2.3.10 evaluate_0.24.0
#> [121] codetools_0.2-20 tibble_3.2.1 cli_3.6.3
#> [124] uwot_0.2.2 arrow_17.0.0.1 xtable_1.8-4
#> [127] reticulate_1.39.0 systemfonts_1.1.0 munsell_0.5.1
#> [130] jquerylib_0.1.4 Rcpp_1.0.13 globals_0.16.3
#> [133] spatstat.random_3.3-2 png_0.1-8 spatstat.univar_3.0-1
#> [136] parallel_4.4.1 assertthat_0.2.1 pkgdown_2.1.0
#> [139] ggplot2_3.5.1 dotCall64_1.2 listenv_0.9.1
#> [142] viridisLite_0.4.2 e1071_1.7-16 scales_1.3.0
#> [145] ggridges_0.5.6 crayon_1.5.3 SeuratObject_5.0.2
#> [148] leiden_0.4.3.1 purrr_1.0.2 rlang_1.1.4
#> [151] cowplot_1.1.3